1 Chapter 3: The Classical DID Design
1.1 Overview
Dataset: Moser & Voena (2012) — moser_voena_didtextbook.dta
The Trading with the Enemy Act (TWEA) allowed US firms to use German-owned patents. Treatment (\(D_{g,t}\)) equals 1 for the 336 subclasses that received German patents, starting in 1919. The dataset contains 289,920 observations (7,248 subclasses × 40 years).
1.2 Panel View
* ssc install panelview, replace
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
panelview patents twea, i(subclass) t(year) type(treat) title("Treatment Status: Moser & Voena (2012)") ylabel(none) ytitle("")
graph export "figures/ch03_panelview_stata.png", replace width(1200)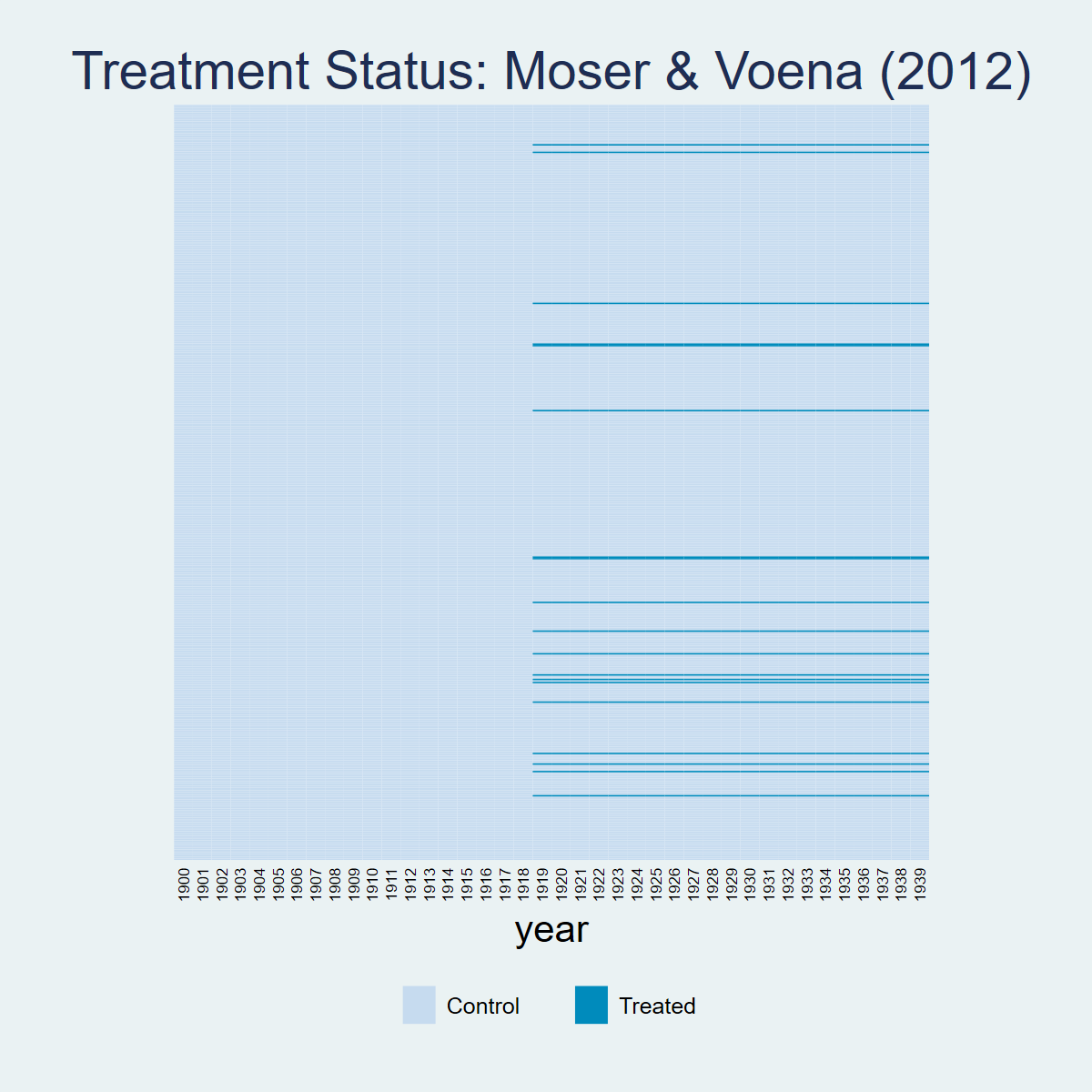
library(panelView)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
png("figures/ch03_panelview_R.png", width = 1000, height = 600)
panelview(patents ~ twea, data = df, index = c("subclass", "year"), type = "treat",
main = "Moser & Voena (2012)", ylab = "")
dev.off()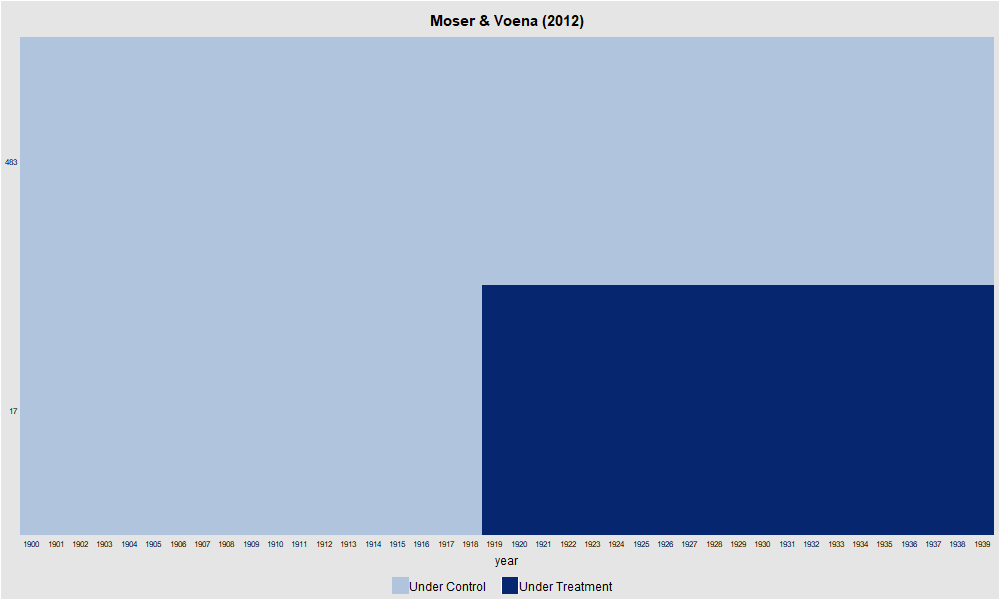
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
from matplotlib.patches import Patch
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
pv = df.pivot_table(index="subclass", columns="year", values="twea", aggfunc="first")
pv_sorted = pv.loc[pv.mean(axis=1).sort_values(ascending=False).index]
cmap = mcolors.ListedColormap(["#D4E6F1", "#00AAAA"])
fig, ax = plt.subplots(figsize=(12, 10))
ax.imshow(pv_sorted.values, aspect="auto", cmap=cmap, interpolation="nearest", vmin=0, vmax=1)
for i in range(0, len(pv_sorted), 10):
ax.axhline(y=i - 0.5, color="white", linewidth=0.15)
ax.set_xticks(range(0, len(pv_sorted.columns), 1))
ax.set_xticklabels([int(c) for c in pv_sorted.columns], rotation=45, ha="right", fontsize=8)
ax.set_yticks([])
ax.set_xlabel("year")
ax.set_title("Treatment Status: Moser & Voena (2012)", fontsize=16)
ax.legend(handles=[Patch(facecolor="#D4E6F1", edgecolor="gray", label="Control"),
Patch(facecolor="#00AAAA", edgecolor="gray", label="Treated")],
loc="lower center", bbox_to_anchor=(0.5, -0.12), ncol=2)
plt.tight_layout()
plt.savefig("figures/ch03_panelview_Python.png", dpi=150, bbox_inches="tight")
plt.show()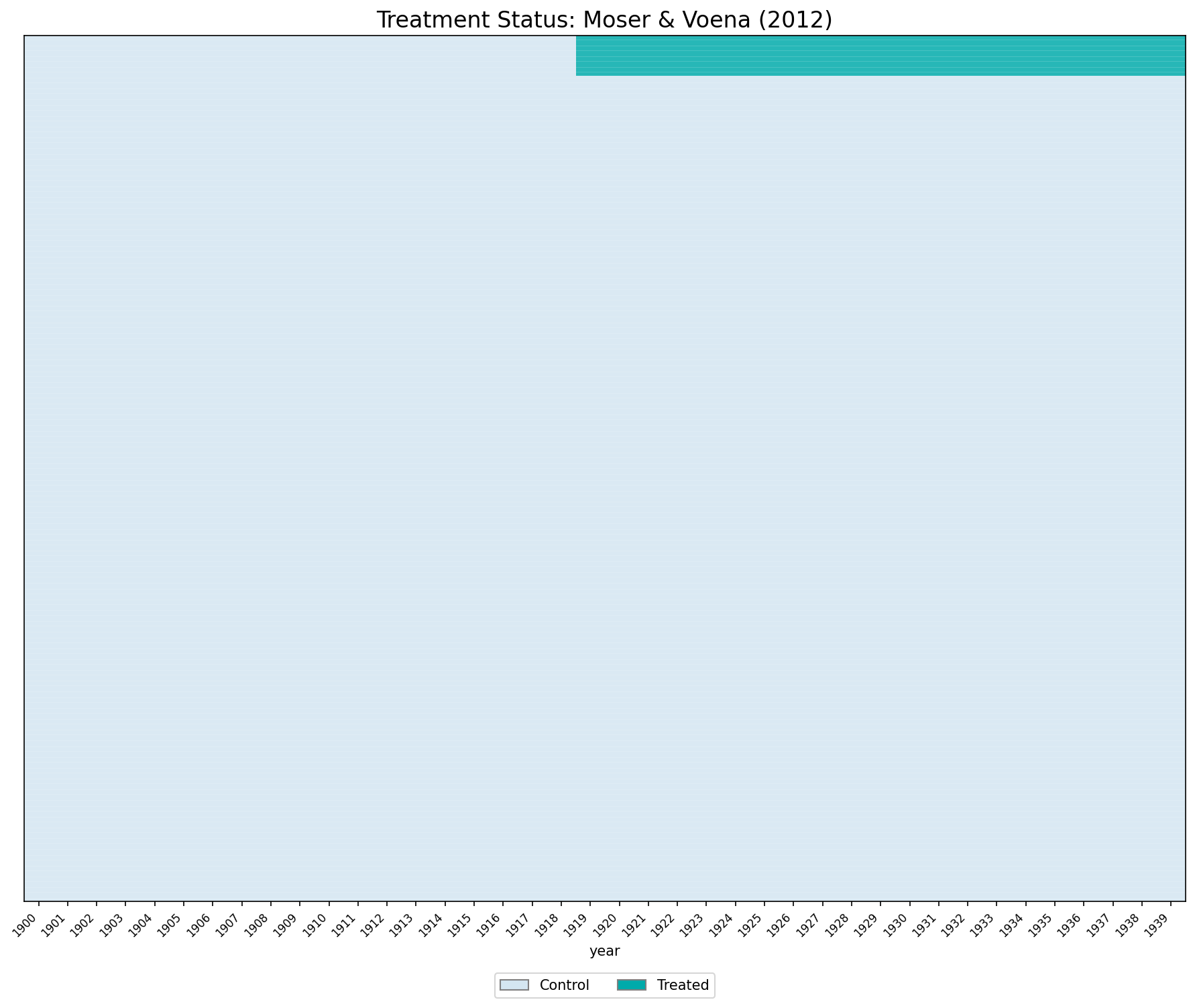
1.3 CQ#1: Static TWFE Regression
Using the
moser_voena_didtextbookdataset, run the TWFE regression of the number of patents in subclass \(g\) and year \(t\) on subclass and year FEs and thetweatreatment, clustering standard errors at the subclass level.
* ssc install reghdfe, replace
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
* Ejemplo 1: xtreg
xtreg patents twea i.year, fe i(subclass) robust cluster(subclass)
* Ejemplo 2: reghdfe
reghdfe patents twea, absorb(subclass year) cluster(subclass)Both commands yield the same coefficient:
(Std. err. adjusted for 7,248 clusters in subclass)
------------------------------------------------------------------------------
| Robust
patents | Coefficient std. err. t P>|t| [95% conf. interval]
-------------+----------------------------------------------------------------
twea | .2882615 .0388871 7.41 0.000 .2120315 .3644915
------------------------------------------------------------------------------library(fixest); library(haven)
options(digits=7)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
# Ejemplo 1
summary(feols(patents ~ twea + i(year) | subclass, data = df, cluster = ~subclass))
# Ejemplo 2
summary(feols(patents ~ twea | subclass + year, data = df, cluster = ~subclass))Ejemplo 1 (feols with i(year)):
Estimate Std. Error t value Pr(>|t|)
twea 0.288262 0.038887 7.412783 1.3770e-13 ***
Ejemplo 2 (feols with subclass + year):
Estimate Std. Error t value Pr(>|t|)
twea 0.288262 0.038887 7.41278 1.377e-13 ***import pandas as pd
import pyfixest as pf
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
# Ejemplo 1
m1 = pf.feols('patents ~ twea + i(year) | subclass', data=df, vcov={'CRV1': 'subclass'})
c1 = m1.coef(); s1 = m1.se(); t1 = m1.tstat(); p1 = m1.pvalue()
for name in c1.index:
print(f"{name:>20s} | {c1[name]:>12.7f} {s1[name]:>10.7f} {t1[name]:>8.2f} {p1[name]:>8.4f}")
# Ejemplo 2
m2 = pf.feols('patents ~ twea | subclass + year', data=df, vcov={'CRV1': 'subclass'})
c2 = m2.coef(); s2 = m2.se(); t2 = m2.tstat(); p2 = m2.pvalue()
print("\n--- Ejemplo 2 ---")
for name in c2.index:
print(f"{name:>20s} | {c2[name]:>12.7f} {s2[name]:>10.7f} {t2[name]:>8.2f} {p2[name]:>8.4f}")Ejemplo 1:
twea | 0.2882615 0.0388871 7.41 0.0000
C(year)[T.1901] | 0.0059327 0.0077529 0.77 0.4442
...
Ejemplo 2:
twea | 0.2882615 0.0388871 7.41 0.0000Result: The coefficient on twea is 0.28826 (\(t = 7.41\), \(p < 0.001\)) across all three languages. Compulsory licensing has a positive and significant effect on US innovation.
1.4 CQ#2: Equivalence between Static TWFE and DID
Verify that \(\hat{\beta}_{fe}\) equals the coefficient on
tweain a regression on the treatment group indicator, the post indicator, andtwea.
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
reg patents treatmentgroup post twea, cluster(subclass) (Std. err. adjusted for 7,248 clusters in subclass)
--------------------------------------------------------------------------------
| Robust
patents | Coefficient std. err. t P>|t| [95% conf. interval]
---------------+----------------------------------------------------------------
treatmentgroup | -.1780874 .0338152 -5.27 0.000 -.2443751 -.1117996
post | .2996666 .0090614 33.07 0.000 .2819036 .3174296
twea | .2882615 .0388846 7.41 0.000 .2120364 .3644867
_cons | .3143656 .0098186 32.02 0.000 .2951183 .3336129
--------------------------------------------------------------------------------library(haven); library(fixest)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
model2 <- feols(patents ~ treatmentgroup + post + twea,
data = df, cluster = ~subclass)
summary(model2)OLS estimation, Dep. Var.: patents
Observations: 289,920
Standard-errors: Clustered (subclass)
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.314366 0.009819 32.01738 < 2.2e-16 ***
treatmentgroup -0.178087 0.033815 -5.26648 1.4306e-07 ***
post 0.299667 0.009061 33.07069 < 2.2e-16 ***
twea 0.288262 0.038885 7.41326 1.3721e-13 ***import pandas as pd
import pyfixest as pf
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
m2 = pf.feols("patents ~ treatmentgroup + post + twea",
data=df, vcov={"CRV1": "subclass"})
print(m2.summary())| Coefficient | Estimate | Std. Error | t value | Pr(>|t|) | 2.5% | 97.5% |
|:---------------|------------|--------------|-----------|------------|---------|---------|
| Intercept | 0.31437 | 0.00982 | 32.01738 | 0.000 | 0.2951 | 0.334 |
| treatmentgroup | -0.17809 | 0.03382 | -5.26648 | 0.000 | -0.2444 | -0.112 |
| post | 0.29967 | 0.00906 | 33.07069 | 0.000 | 0.2819 | 0.317 |
| twea | 0.28826 | 0.03889 | 7.41326 | 0.000 | 0.2120 | 0.365 |Result: The coefficient on twea is identical (0.28826) in both specifications across all three languages.
1.5 CQ#3: Testing Randomized Treatment
Regress patents on the treatment group indicator, restricting the sample to years before 1919.
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
reg patents treatmentgroup if year<=1918, cluster(subclass) (Std. err. adjusted for 7,248 clusters in subclass)
--------------------------------------------------------------------------------
| Robust
patents | Coefficient std. err. t P>|t| [95% conf. interval]
---------------+----------------------------------------------------------------
treatmentgroup | -.1780874 .0338152 -5.27 0.000 -.244375 -.1117997
_cons | .3143656 .0098186 32.02 0.000 .2951183 .3336128
--------------------------------------------------------------------------------library(haven); library(fixest); library(dplyr)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
model3 <- feols(patents ~ treatmentgroup,
data = df %>% filter(year <= 1918), cluster = ~subclass)
summary(model3)OLS estimation, Dep. Var.: patents
Observations: 137,712
Standard-errors: Clustered (subclass)
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.314366 0.009819 32.01743 < 2.2e-16 ***
treatmentgroup -0.178087 0.033815 -5.26649 1.4306e-07 ***import pandas as pd
import pyfixest as pf
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
df_pre = df[df["year"] <= 1918].copy()
m3 = pf.feols("patents ~ treatmentgroup",
data=df_pre, vcov={"CRV1": "subclass"})
print(m3.summary())| Coefficient | Estimate | Std. Error | t value | Pr(>|t|) | 2.5% | 97.5% |
|:---------------|------------|--------------|-----------|------------|---------|---------|
| Intercept | 0.31437 | 0.00982 | 32.01743 | 0.000 | 0.2951 | 0.334 |
| treatmentgroup | -0.17809 | 0.03382 | -5.26649 | 0.000 | -0.2444 | -0.112 |Result: The coefficient is −0.17809 (\(t = -5.27\), \(p < 0.001\)). Treatment is not randomly assigned — treated subclasses had fewer patents before the TWEA.
1.6 CQ#4–7: Event-Study TWFE Regression
Estimate the event-study TWFE regression. Test whether pre-trend coefficients are jointly significant. Verify that \(\hat{\beta}_1^{fe}\) equals the DID from equation (3.7).
The event-study dummies reltimeminus1–reltimeminus18 and reltimeplus1–reltimeplus21 are already included in the dataset. The reference period is 1918 (the last pre-treatment year).
Now estimate the event-study TWFE regression:
Now estimate the event-study TWFE regression:
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
forvalues l = 1/18 {
gen reltimeminus`l' = treatmentgroup * (year == 1918 - `l')
}
forvalues l = 1/21 {
gen reltimeplus`l' = treatmentgroup * (year == 1918 + `l')
}
reg patents i.year treatmentgroup reltimeminus* reltimeplus*, cluster(subclass)
* F-test on pre-trends
test reltimeminus1 reltimeminus2 reltimeminus3 reltimeminus4 reltimeminus5 reltimeminus6 reltimeminus7 reltimeminus8 reltimeminus9 reltimeminus10 reltimeminus11 reltimeminus12 reltimeminus13 reltimeminus14 reltimeminus15 reltimeminus16 reltimeminus17 reltimeminus18 F( 18, 7247) = 3.79
Prob > F = 0.0000* Verify equation (3.7) for l=1 (continues from above)
quietly summarize patents if year==1919 & treatmentgroup==1
scalar m1 = r(mean)
quietly summarize patents if year==1918 & treatmentgroup==1
scalar m2 = r(mean)
quietly summarize patents if year==1919 & treatmentgroup==0
scalar m3 = r(mean)
quietly summarize patents if year==1918 & treatmentgroup==0
scalar m4 = r(mean)
display "DID manual = " m1-m2-(m3-m4)DID manual (eq 3.7, l=1) = .02352017library(haven); library(fixest); library(car)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
for (l in 1:18) df[[paste0("reltimeminus", l)]] <- as.numeric(df$treatmentgroup == 1 & df$year == 1918 - l)
for (l in 1:21) df[[paste0("reltimeplus", l)]] <- as.numeric(df$treatmentgroup == 1 & df$year == 1918 + l)
event_formula <- as.formula(paste(
"patents ~ i(year) + treatmentgroup +",
paste(paste0("reltimeminus", 1:18), collapse = " + "), "+",
paste(paste0("reltimeplus", 1:21), collapse = " + ")))
model4 <- feols(event_formula, data = df, cluster = ~subclass)
# F-test on pre-trends
library(car)
linearHypothesis(model4, paste0("reltimeminus", 1:18))
# DID manual
m1 <- mean(df$patents[df$year == 1919 & df$treatmentgroup == 1])
m2 <- mean(df$patents[df$year == 1918 & df$treatmentgroup == 1])
m3 <- mean(df$patents[df$year == 1919 & df$treatmentgroup == 0])
m4 <- mean(df$patents[df$year == 1918 & df$treatmentgroup == 0])
cat("DID manual:", m1 - m2 - (m3 - m4))Chisq = 68.25, p-value = 8.915e-08
DID manual (eq 3.7, l=1): 0.02352017
reltimeplus1 coef: 0.02352017import pandas as pd
import pyfixest as pf
import numpy as np
from scipy import stats
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
for l in range(1, 19):
df[f"reltimeminus{l}"] = (df["treatmentgroup"] * (df["year"] == 1918 - l)).astype(float)
for l in range(1, 22):
df[f"reltimeplus{l}"] = (df["treatmentgroup"] * (df["year"] == 1918 + l)).astype(float)
minus_vars = [f"reltimeminus{i}" for i in range(1, 19)]
plus_vars = [f"reltimeplus{i}" for i in range(1, 22)]
all_reltime = minus_vars + plus_vars
fml_es = "patents ~ " + " + ".join(all_reltime) + " + treatmentgroup | year"
m4 = pf.feols(fml_es, data=df, vcov={"CRV1": "subclass"})
# F-test on pre-trends
c = m4.coef()
v_mat = pd.DataFrame(m4._vcov, index=c.index, columns=c.index)
pre_names = [x for x in minus_vars if x in c.index]
beta_pre = c[pre_names].values
V_pre = v_mat.loc[pre_names, pre_names].values
F_stat = beta_pre @ np.linalg.inv(V_pre) @ beta_pre / len(pre_names)
p_val = 1 - stats.f.cdf(F_stat, len(pre_names), df["subclass"].nunique() - 1)
print(f"F-test: F({len(pre_names)}, {df['subclass'].nunique()-1}) = {F_stat:.4f}, p = {p_val:.4f}")
# DID manual
m1v = df.loc[(df["year"]==1919) & (df["treatmentgroup"]==1), "patents"].mean()
m2v = df.loc[(df["year"]==1918) & (df["treatmentgroup"]==1), "patents"].mean()
m3v = df.loc[(df["year"]==1919) & (df["treatmentgroup"]==0), "patents"].mean()
m4v = df.loc[(df["year"]==1918) & (df["treatmentgroup"]==0), "patents"].mean()
print(f"DID manual (eq 3.7, l=1): {m1v - m2v - (m3v - m4v):.8f}")
print(f"reltimeplus1 coef: {m4.coef()['reltimeplus1']:.8f}")F-test: F(18, 7247) = 3.7917, p = 0.0000
DID manual (eq 3.7, l=1): 0.02352017
reltimeplus1 coef: 0.02352017Result: Pre-trends are jointly significant (\(F = 3.79\), \(p < 0.001\)). The manual DID (0.02352) matches reltimeplus1 exactly.
1.6.1 Figure 3.2: TWFE Event-Study Plot
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
reg patents i.year treatmentgroup reltimeminus* reltimeplus*, cluster(subclass)
coefplot, keep(reltimeminus* reltimeplus*) vertical yline(0) xline(18.5, lpattern(dash)) xtitle("Relative time to year before TWEA") ytitle("Effect") title("TWFE Event-study estimates") ciopts(recast(rcap) color(cranberry))
graph export "figures/ch03_fig32_es_twfe.png", replace width(1200)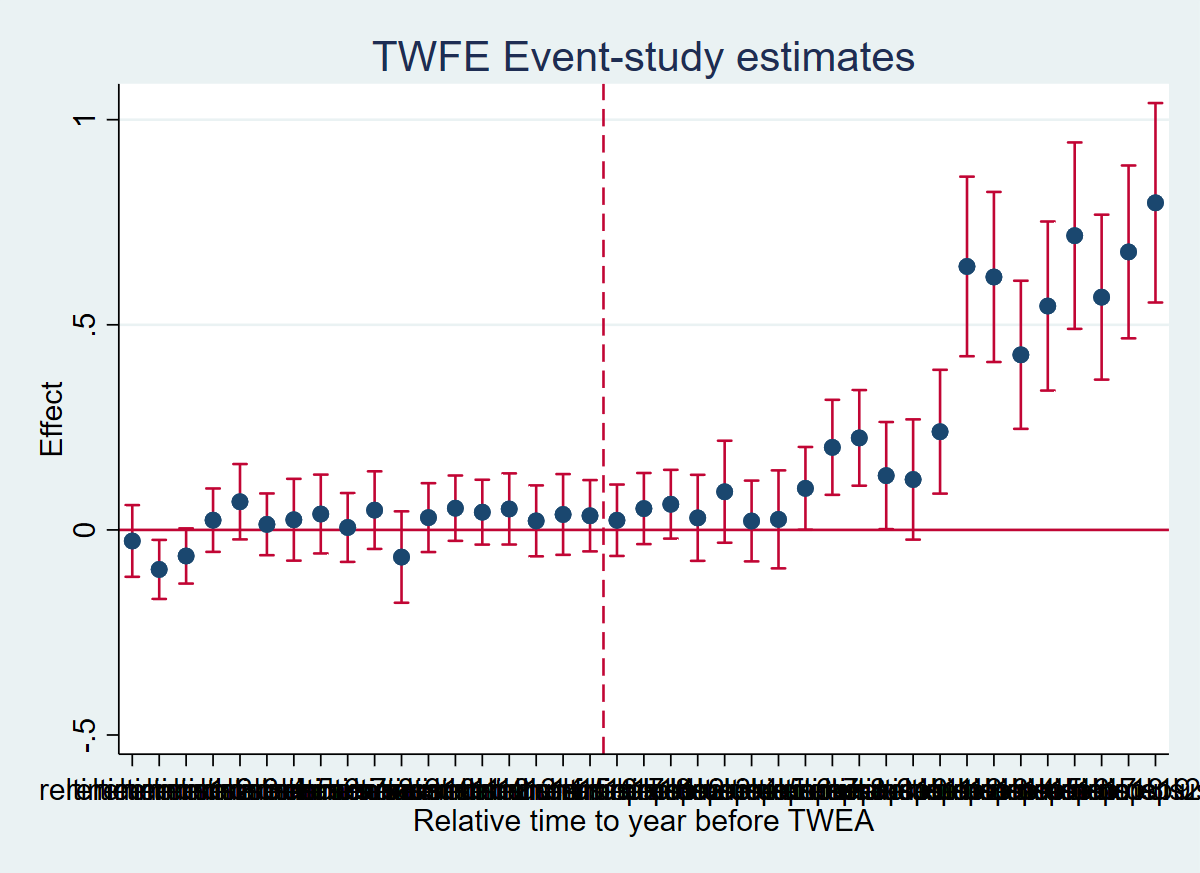
library(fixest); library(haven)
options(digits=7)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
reltimeminus_vars <- paste0("reltimeminus", 1:18)
reltimeplus_vars <- paste0("reltimeplus", 1:21)
formula_str <- paste("patents ~", paste(reltimeminus_vars, collapse = " + "), "+",
paste(reltimeplus_vars, collapse = " + "), "+ treatmentgroup | year")
model4 <- feols(as.formula(formula_str), data = df, cluster = ~subclass)
rt <- c(-(18:1), 0, 1:21)
coefs <- c(rev(coef(model4)[paste0("reltimeminus", 18:1)]), 0,
coef(model4)[paste0("reltimeplus", 1:21)])
ses <- c(rev(se(model4)[paste0("reltimeminus", 18:1)]), 0,
se(model4)[paste0("reltimeplus", 1:21)])
png("figures/ch03_fig32_es_twfe_R.png", width = 1000, height = 600)
plot(rt, coefs, type = "b", pch = 19, col = "navy", cex = 0.6,
xlab = "Relative time to year before TWEA", ylab = "Effect",
main = "TWFE Event-study estimates", ylim = range(coefs - 1.96*ses, coefs + 1.96*ses))
arrows(rt, coefs - 1.96*ses, rt, coefs + 1.96*ses,
length = 0.03, angle = 90, code = 3, col = "firebrick")
abline(h = 0, col = "black", lwd = 0.8)
abline(v = 0, col = "black", lty = 2)
dev.off()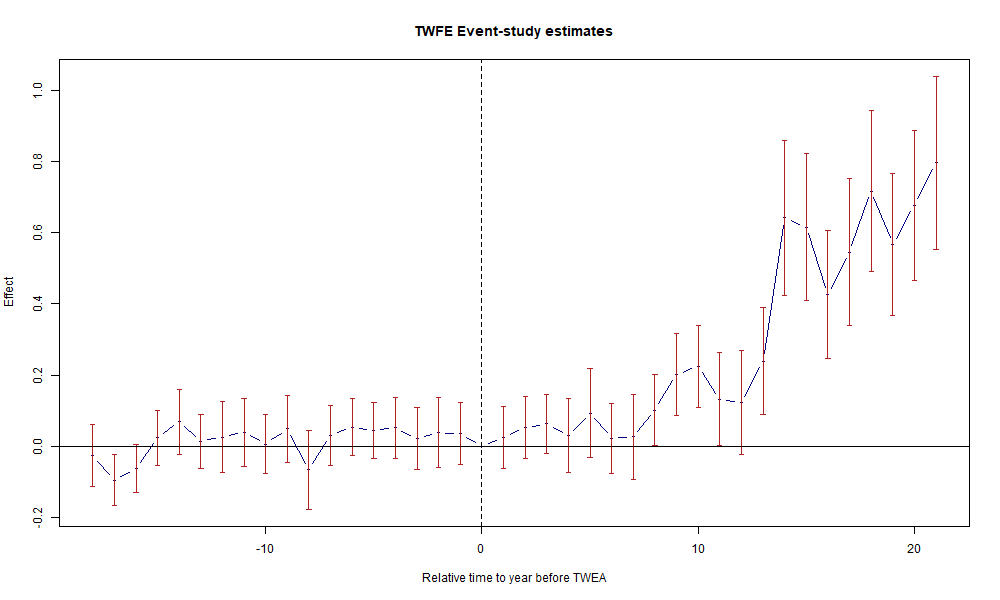
import pandas as pd
import pyfixest as pf
import matplotlib.pyplot as plt
import numpy as np
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
minus_vars = [f"reltimeminus{i}" for i in range(1, 19)]
plus_vars = [f"reltimeplus{i}" for i in range(1, 22)]
fml = "patents ~ " + " + ".join(minus_vars + plus_vars) + " + treatmentgroup | year"
m4 = pf.feols(fml, data=df, vcov={'CRV1': 'subclass'})
rt = list(range(-18, 0)) + [0] + list(range(1, 22))
c4 = [m4.coef()[f"reltimeminus{i}"] for i in range(18, 0, -1)] + [0] + \
[m4.coef()[f"reltimeplus{i}"] for i in range(1, 22)]
s4 = [m4.se()[f"reltimeminus{i}"] for i in range(18, 0, -1)] + [0] + \
[m4.se()[f"reltimeplus{i}"] for i in range(1, 22)]
c4, s4 = np.array(c4), np.array(s4)
fig, ax = plt.subplots(figsize=(10, 6))
ax.plot(rt, c4, 'o-', color='navy', markersize=4, linewidth=1.2)
ax.errorbar(rt, c4, yerr=1.96*s4, fmt='none', ecolor='firebrick', capsize=2, linewidth=1)
ax.axhline(0, color='black', linewidth=0.8)
ax.axvline(0, color='black', linestyle='--', linewidth=0.8)
ax.set_xlabel("Relative time to year before TWEA")
ax.set_ylabel("Effect")
ax.set_title("TWFE Event-study estimates")
plt.tight_layout()
plt.savefig("figures/ch03_fig32_es_twfe_Python.png", dpi=150, bbox_inches="tight")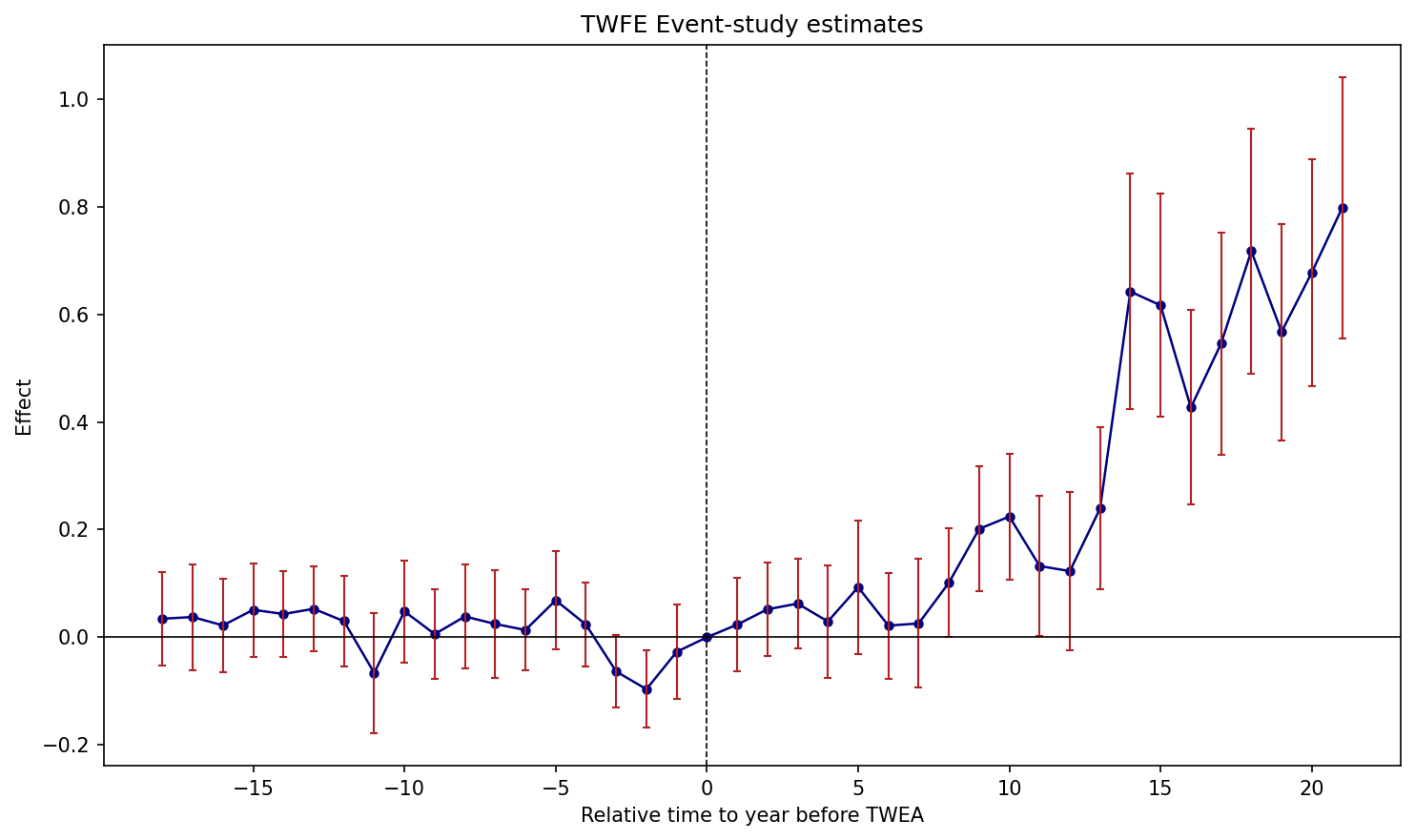
Figure 3.2: Effects of compulsory licensing on US innovation, according to an ES-TWFER. Standard errors clustered at the patent subclass level. 95% confidence intervals shown in red.
1.7 CQ#9: Event-Study without Pre-Trends
Re-estimate the event-study using only post-treatment dummies. This is numerically equivalent to Borusyak, Jaravel, and Spiess (2024).
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
forvalues l = 1/21 {
gen reltimeplus`l' = treatmentgroup * (year == 1918 + `l')
}
reg patents i.yearpost treatmentgroup reltimeplus*, cluster(subclass) (Std. err. adjusted for 7,248 clusters in subclass)
--------------------------------------------------------------------------------
| Robust
patents | Coefficient std. err. t P>|t| [95% conf. interval]
---------------+----------------------------------------------------------------
treatmentgroup | -.1780874 .0338176 -5.27 0.000 -.2443797 -.1117951
reltimeplus1 | .0108007 .0309763 0.35 0.727 -.049922 .0715233
reltimeplus2 | .0393018 .0302047 1.30 0.193 -.0199081 .0985118
reltimeplus3 | .0498425 .0359726 1.39 0.166 -.0206743 .1203593
reltimeplus4 | .0166497 .0418098 0.40 0.690 -.0653098 .0986092
reltimeplus5 | .0803071 .049547 1.62 0.105 -.0168195 .1774337
reltimeplus6 | .0089819 .041055 0.22 0.827 -.0714978 .0894616
reltimeplus7 | .0129501 .0472633 0.27 0.784 -.0796997 .1056
reltimeplus8 | .0885536 .0432021 2.05 0.040 .0038648 .1732424
reltimeplus9 | .1885454 .0479442 3.93 0.000 .0945607 .28253
reltimeplus10 | .2115488 .050361 4.20 0.000 .1128266 .310271
reltimeplus11 | .1196796 .0520212 2.30 0.021 .017703 .2216563
reltimeplus12 | .1101517 .0593023 1.86 0.063 -.0060982 .2264015
reltimeplus13 | .2266778 .0691283 3.28 0.001 .0911663 .3621893
reltimeplus14 | .6294969 .1040565 6.05 0.000 .4255158 .833478
reltimeplus15 | .6038686 .0994844 6.07 0.000 .4088502 .798887
reltimeplus16 | .4141158 .0858483 4.82 0.000 .2458282 .5824034
reltimeplus17 | .5330394 .0957086 5.57 0.000 .3454227 .7206561
reltimeplus18 | .7046044 .109204 6.45 0.000 .4905328 .918676
reltimeplus19 | .5546581 .0930222 5.96 0.000 .3723075 .7370087
reltimeplus20 | .6650045 .1024705 6.49 0.000 .4641324 .8658766
reltimeplus21 | .7847135 .1190138 6.59 0.000 .5514119 1.018015
_cons | .3143656 .0098193 32.02 0.000 .2951169 .3336142
--------------------------------------------------------------------------------library(haven); library(fixest)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
for (l in 1:21) df[[paste0("reltimeplus", l)]] <- as.numeric(df$treatmentgroup == 1 & df$year == 1918 + l)
post_formula <- as.formula(paste(
"patents ~ i(yearpost) + treatmentgroup +",
paste(paste0("reltimeplus", 1:21), collapse = " + ")))
model5 <- feols(post_formula, data = df, cluster = ~subclass)
summary(model5) Estimate Std. Error t value Pr(>|t|)
treatmentgroup -0.178087 0.033818 -5.266118 1.4334e-07 ***
reltimeplus1 0.010801 0.030976 0.348674 7.2734e-01
reltimeplus2 0.039302 0.030205 1.301183 1.9324e-01
reltimeplus3 0.049843 0.035973 1.385634 1.6590e-01
reltimeplus4 0.016650 0.041810 0.398234 6.9050e-01
reltimeplus5 0.080307 0.049547 1.620753 1.0514e-01
reltimeplus6 0.008982 0.041055 0.218776 8.2685e-01
reltimeplus7 0.012950 0.047263 0.274032 7.8404e-01
reltimeplus8 0.088554 0.043202 2.049798 4.0405e-02 *
reltimeplus9 0.188545 0.047944 3.932590 8.4570e-05 ***
reltimeplus10 0.211549 0.050361 4.201191 2.6755e-05 ***
reltimeplus11 0.119680 0.052021 2.300631 2.1441e-02 *
reltimeplus12 0.110152 0.059302 1.857479 6.3256e-02
reltimeplus13 0.226678 0.069128 3.278641 1.0478e-03 **
reltimeplus14 0.629497 0.104057 6.050261 1.5073e-09 ***
reltimeplus15 0.603869 0.099484 6.069422 1.3510e-09 ***
reltimeplus16 0.414116 0.085848 4.823560 1.4305e-06 ***
reltimeplus17 0.533039 0.095709 5.568517 2.6464e-08 ***
reltimeplus18 0.704604 0.109204 6.452210 1.1588e-10 ***
reltimeplus19 0.554658 0.093022 5.963752 2.5802e-09 ***
reltimeplus20 0.665005 0.102471 6.490150 9.2071e-11 ***
reltimeplus21 0.784714 0.119014 6.593469 4.5985e-11 ***
---
RMSE: 1.33696 Adj. R2: 0.025481import pandas as pd
import pyfixest as pf
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
for l in range(1, 22):
df[f"reltimeplus{l}"] = (df["treatmentgroup"] * (df["year"] == 1918 + l)).astype(float)
plus_vars = [f"reltimeplus{i}" for i in range(1, 22)]
fml5 = "patents ~ " + " + ".join(plus_vars) + " + treatmentgroup | yearpost"
m5 = pf.feols(fml5, data=df, vcov={"CRV1": "subclass"})
print(m5.summary())| Coefficient | Estimate | Std. Error | t value | Pr(>|t|) | 2.5% | 97.5% |
|:---------------|------------|--------------|-----------|------------|---------|---------|
| reltimeplus1 | 0.01080 | 0.03098 | 0.34867 | 0.72734 | -0.0499 | 0.072 |
| reltimeplus2 | 0.03930 | 0.03020 | 1.30118 | 0.19324 | -0.0199 | 0.099 |
| reltimeplus3 | 0.04984 | 0.03597 | 1.38563 | 0.16590 | -0.0207 | 0.120 |
| reltimeplus4 | 0.01665 | 0.04181 | 0.39823 | 0.69050 | -0.0653 | 0.099 |
| reltimeplus5 | 0.08031 | 0.04955 | 1.62075 | 0.10514 | -0.0168 | 0.177 |
| reltimeplus6 | 0.00898 | 0.04106 | 0.21878 | 0.82685 | -0.0715 | 0.089 |
| reltimeplus7 | 0.01295 | 0.04726 | 0.27403 | 0.78404 | -0.0797 | 0.106 |
| reltimeplus8 | 0.08855 | 0.04320 | 2.04980 | 0.04041 | 0.0039 | 0.173 |
| reltimeplus9 | 0.18855 | 0.04794 | 3.93259 | 0.00008 | 0.0946 | 0.283 |
| reltimeplus10 | 0.21155 | 0.05036 | 4.20119 | 0.00003 | 0.1128 | 0.310 |
| reltimeplus11 | 0.11968 | 0.05202 | 2.30063 | 0.02144 | 0.0177 | 0.222 |
| reltimeplus12 | 0.11015 | 0.05930 | 1.85748 | 0.06326 | -0.0061 | 0.226 |
| reltimeplus13 | 0.22668 | 0.06913 | 3.27864 | 0.00105 | 0.0912 | 0.362 |
| reltimeplus14 | 0.62950 | 0.10406 | 6.05026 | 0.00000 | 0.4255 | 0.833 |
| reltimeplus15 | 0.60387 | 0.09948 | 6.06942 | 0.00000 | 0.4089 | 0.799 |
| reltimeplus16 | 0.41412 | 0.08585 | 4.82356 | 0.00000 | 0.2458 | 0.582 |
| reltimeplus17 | 0.53304 | 0.09571 | 5.56852 | 0.00000 | 0.3454 | 0.721 |
| reltimeplus18 | 0.70460 | 0.10920 | 6.45221 | 0.00000 | 0.4905 | 0.919 |
| reltimeplus19 | 0.55466 | 0.09302 | 5.96375 | 0.00000 | 0.3723 | 0.737 |
| reltimeplus20 | 0.66500 | 0.10247 | 6.49015 | 0.00000 | 0.4641 | 0.866 |
| reltimeplus21 | 0.78471 | 0.11901 | 6.59347 | 0.00000 | 0.5514 | 1.018 |
| treatmentgroup | -0.17809 | 0.03382 | -5.26612 | 0.00000 | -0.2444 | -0.112 |
---
RMSE: 1.337 R2: 0.026 R2 Within: 0.0011.7.1 Figure 3.5: Comparison of two ES-TWFER estimators
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
reg patents i.year treatmentgroup reltimeminus* reltimeplus*, cluster(subclass)
estimates store m1
reg patents i.yearpost treatmentgroup reltimeplus*, cluster(subclass)
estimates store m2
coefplot (m1, label("With Pre-Periods") mcolor(gs10) ciopts(lcolor(gs10)) connect(l) lcolor(gs10) msymbol(O) msize(small)) (m2, label("Without Pre-Periods") mcolor(gs4) ciopts(lcolor(gs4)) connect(l) lcolor(gs4) msymbol(O) msize(small)), keep(reltimeplus*) vertical yline(0, lcolor(gs12) lpattern(solid)) title("TWFE Event-study estimates") xtitle("Relative time to year before TWEA") ytitle("Effect") legend(rows(1) position(6)) scheme(s2mono) rename(reltimeplus1="0" reltimeplus4="3" reltimeplus7="6" reltimeplus10="9" reltimeplus13="12" reltimeplus16="15" reltimeplus19="18" reltimeplus21="21" reltimeplus2=" " reltimeplus3=" " reltimeplus5=" " reltimeplus6=" " reltimeplus8=" " reltimeplus9=" " reltimeplus11=" " reltimeplus12=" " reltimeplus14=" " reltimeplus15=" " reltimeplus17=" " reltimeplus18=" " reltimeplus20=" ")
graph export "figures/ch03_fig35_comparison.png", replace width(1200)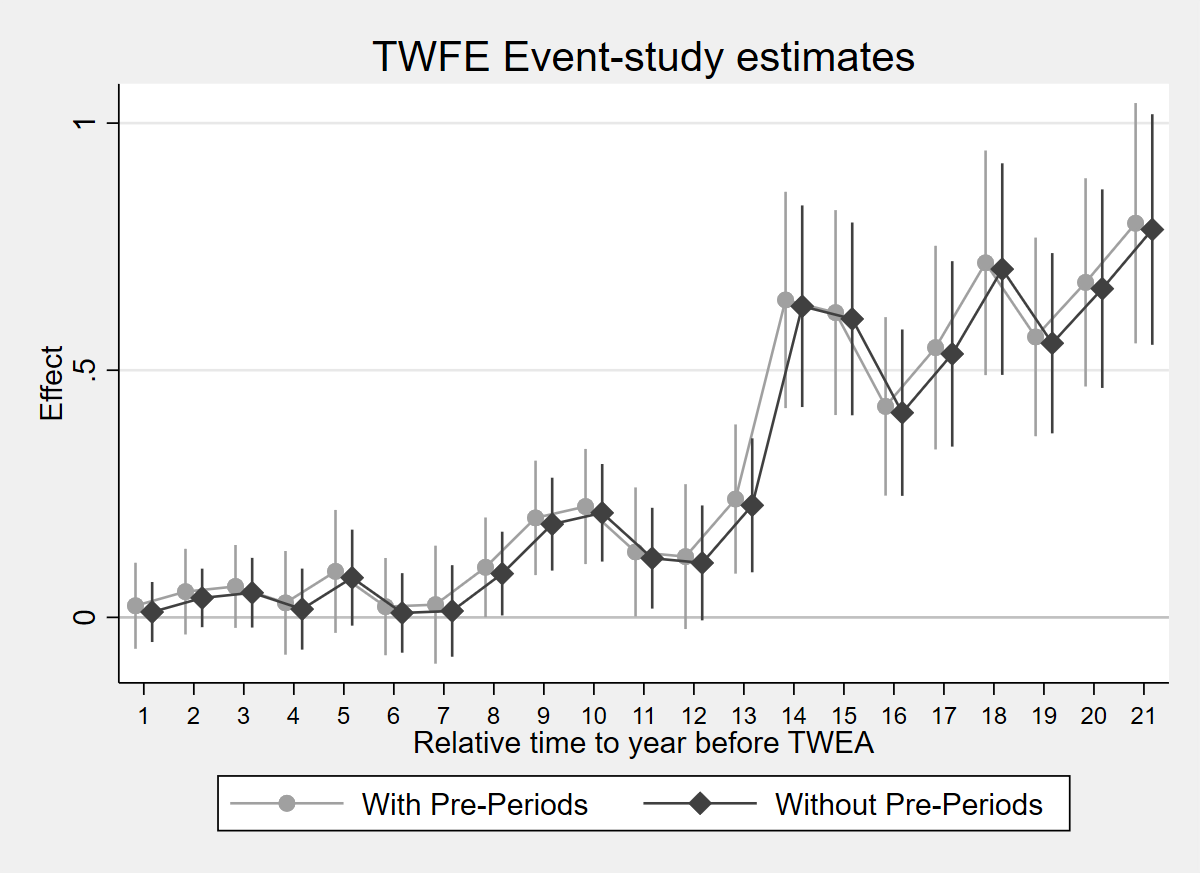
library(fixest); library(dplyr); library(ggplot2)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
reltimeminus_vars <- paste0("reltimeminus", 1:18)
reltimeplus_vars <- paste0("reltimeplus", 1:21)
fml1 <- as.formula(paste("patents ~ i(year) + treatmentgroup +", paste(c(reltimeminus_vars, reltimeplus_vars), collapse = " + ")))
m1 <- feols(fml1, data = df, cluster = ~subclass)
fml2 <- as.formula(paste("patents ~ i(yearpost) + treatmentgroup +", paste(reltimeplus_vars, collapse = " + ")))
m2 <- feols(fml2, data = df, cluster = ~subclass)
coef1 <- data.frame(x = 1:21, est = coef(m1)[reltimeplus_vars], se = se(m1)[reltimeplus_vars], model = "With Pre-Periods")
coef2 <- data.frame(x = 1:21, est = coef(m2)[reltimeplus_vars], se = se(m2)[reltimeplus_vars], model = "Without Pre-Periods")
plotdf <- rbind(coef1, coef2)
plotdf$lo <- plotdf$est - 1.96 * plotdf$se
plotdf$hi <- plotdf$est + 1.96 * plotdf$se
ggplot(plotdf, aes(x = x, y = est, color = model)) +
geom_line(position = position_dodge(0.3)) +
geom_point(position = position_dodge(0.3), size = 1.5) +
geom_errorbar(aes(ymin = lo, ymax = hi), width = 0.3, position = position_dodge(0.3)) +
geom_hline(yintercept = 0, linetype = "solid", color = "grey70") +
scale_color_manual(values = c("With Pre-Periods" = "grey70", "Without Pre-Periods" = "grey30")) +
scale_x_continuous(breaks = c(1, 4, 7, 10, 13, 16, 19, 21), labels = c("0", "3", "6", "9", "12", "15", "18", "21")) +
labs(x = "Relative time to year before TWEA", y = "Effect", title = "TWFE Event-study estimates", color = "") +
theme_minimal() + theme(legend.position = "bottom")
ggsave("figures/ch03_fig35_comparison_R.png", width = 8, height = 5, dpi = 150)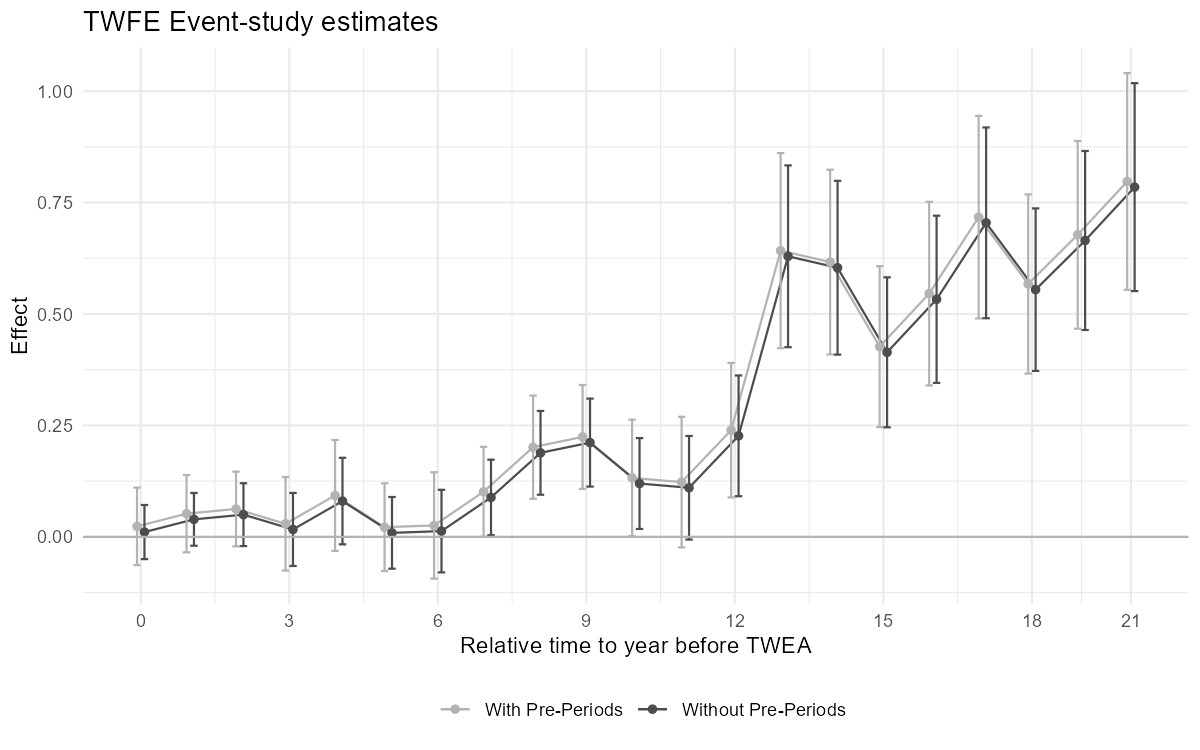
import pandas as pd
import numpy as np
import pyfixest as pf
import matplotlib.pyplot as plt
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
minus_vars = [f"reltimeminus{i}" for i in range(1, 19)]
plus_vars = [f"reltimeplus{i}" for i in range(1, 22)]
fml1 = "patents ~ " + " + ".join(minus_vars + plus_vars) + " + C(year) + treatmentgroup"
m1 = pf.feols(fml1, data=df, vcov={"CRV1": "subclass"})
fml2 = "patents ~ " + " + ".join(plus_vars) + " + C(yearpost) + treatmentgroup"
m2 = pf.feols(fml2, data=df, vcov={"CRV1": "subclass"})
x = np.arange(1, 22)
c1 = np.array([m1.coef()[v] for v in plus_vars])
s1 = np.array([m1.se()[v] for v in plus_vars])
c2 = np.array([m2.coef()[v] for v in plus_vars])
s2 = np.array([m2.se()[v] for v in plus_vars])
fig, ax = plt.subplots(figsize=(8, 5))
ax.errorbar(x - 0.1, c1, yerr=1.96*s1, fmt='-o', color='grey', markersize=3, linewidth=1, capsize=2, capthick=0.8, elinewidth=0.8, label='With Pre-Periods')
ax.errorbar(x + 0.1, c2, yerr=1.96*s2, fmt='-o', color='black', markersize=3, linewidth=1, capsize=2, capthick=0.8, elinewidth=0.8, label='Without Pre-Periods')
ax.axhline(y=0, color='grey', linestyle='-', linewidth=0.5)
ax.set_xticks([1, 4, 7, 10, 13, 16, 19, 21])
ax.set_xticklabels(["0", "3", "6", "9", "12", "15", "18", "21"])
ax.set_xlabel("Relative time to year before TWEA")
ax.set_ylabel("Effect")
ax.set_title("TWFE Event-study estimates")
ax.legend(loc='upper left', frameon=False)
ax.set_ylim(-0.25, 1.1)
plt.tight_layout()
plt.savefig("figures/ch03_fig35_comparison_Python.png", dpi=150)
plt.show()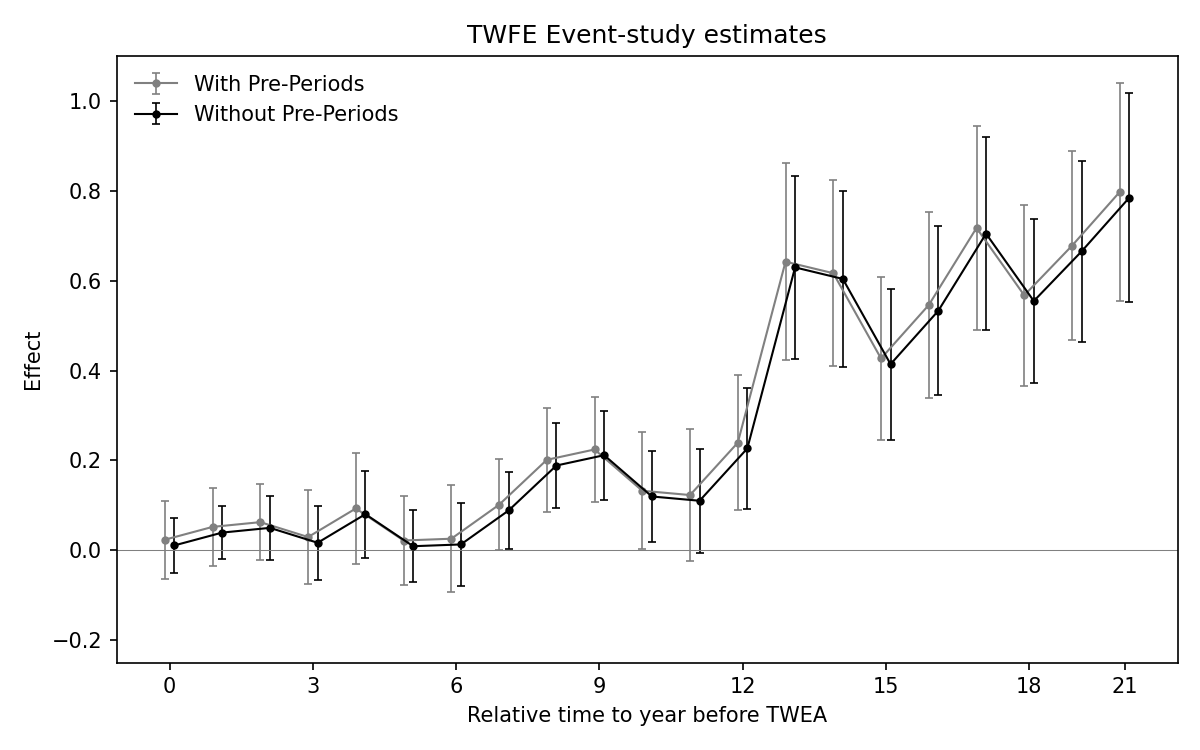
Figure 3.5: Effects of compulsory licensing on US innovation: comparison of two ES-TWFER estimators. The coefficients shown in grey come from the usual ES-TWFER with FEs for pre-treatment periods. The coefficients shown in black come from the imputation ES-TWFER without FEs for pre-treatment periods. Standard errors are clustered at the patent subclass level. 95% confidence intervals shown.
1.8 CQ#8: Linear Pre-Trends We Could Fail to Detect
Use the
pretrendspackage (Roth, 2022) to compute the slope detectable with 50% power using 6 pre-periods.
* ssc install pretrends, replace
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
forvalues l = 1/18 {
gen reltimeminus`l' = treatmentgroup * (year == 1918 - `l')
}
forvalues l = 1/21 {
gen reltimeplus`l' = treatmentgroup * (year == 1918 + `l')
}
reghdfe patents reltimeminus* reltimeplus*, absorb(treatmentgroup year) cluster(subclass)
pretrends power 0.5, numpre(6)Slope for 50% power = .0098235library(haven); library(fixest); library(pretrends)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
for (l in 1:18) df[[paste0("reltimeminus", l)]] <- as.numeric(df$treatmentgroup == 1 & df$year == 1918 - l)
for (l in 1:21) df[[paste0("reltimeplus", l)]] <- as.numeric(df$treatmentgroup == 1 & df$year == 1918 + l)
model6 <- feols(as.formula(paste(
"patents ~", paste(paste0("reltimeminus", 1:18), collapse = " + "), "+",
paste(paste0("reltimeplus", 1:21), collapse = " + "), "| treatmentgroup + year")),
data = df, cluster = ~subclass)
# Extract coefficients and vcov for first 6 pre-periods
pretrend_coefs_6 <- paste0("reltimeminus", 1:6)
beta <- coef(model6)[pretrend_coefs_6]
sigma <- vcov(model6)[pretrend_coefs_6, pretrend_coefs_6]
slope50 <- slope_for_power(sigma = sigma, targetPower = 0.5,
tVec = -6:-1, referencePeriod = 0)
cat("Slope for 50% power:", slope50)Slope for 50% power (numpre=6): 0.009796021import pandas as pd
import pyfixest as pf
from pretrends import slope_for_power
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
for l in range(1, 19):
df[f"reltimeminus{l}"] = (df["treatmentgroup"] * (df["year"] == 1918 - l)).astype(float)
for l in range(1, 22):
df[f"reltimeplus{l}"] = (df["treatmentgroup"] * (df["year"] == 1918 + l)).astype(float)
minus_vars = [f"reltimeminus{i}" for i in range(1, 19)]
plus_vars = [f"reltimeplus{i}" for i in range(1, 22)]
# Full model (matches Stata: reghdfe patents reltimeminus* reltimeplus*)
fml6 = "patents ~ " + " + ".join(minus_vars + plus_vars) + " | treatmentgroup + year"
m6 = pf.feols(fml6, data=df, vcov={"CRV1": "subclass"})
# Extract vcov for 6 closest pre-periods
pre6 = [f"reltimeminus{i}" for i in range(1, 7)]
vcov6 = pd.DataFrame(m6._vcov, index=m6.coef().index,
columns=m6.coef().index).loc[pre6, pre6].values
slope50 = slope_for_power(vcov6, target_power=0.5,
t_vec=[-6, -5, -4, -3, -2, -1])
print(f"Slope for 50% power (numpre=6): {slope50:.7f}")Slope for 50% power (numpre=6): 0.0098252Note: All three platforms give essentially identical results: Stata ≈ 0.0098235, R ≈ 0.009796, Python ≈ 0.0098252. The Python pretrends package (based on Roth 2022) uses the exact multivariate normal CDF, matching Stata and R.
1.9 CQ#10: Variance of Treatment Effects
Compare the variance of
diffpatentswrt1918in treatment and control groups in 1932.
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
sdtest diffpatentswrt1918 if year==1932, by(treatmentgroup)
scalar sd_effects = r(sd_2) - r(sd_1)
reg diffpatentswrt1918 treatmentgroup if year==1932
display "CI: [" _b[treatmentgroup] - 1.96*sd_effects ", " _b[treatmentgroup] + 1.96*sd_effects "]"Variance ratio test
--------------------------------------------------------------
Group | Obs Std. dev.
---------+------------------------
0 | 6,912 1.772807
1 | 336 2.011905
--------------------------------------------------------------
ratio = sd(0) / sd(1) f = 0.7764
Pr(F < f) = 0.0004 2*Pr(F < f) = 0.0008
sd_2 - sd_1 = .23909779
beta = .64221644
CI: [.17358476, 1.1108481]library(haven); library(dplyr)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
df_1932 <- df %>% filter(year == 1932)
var_test <- var.test(diffpatentswrt1918 ~ treatmentgroup, data = df_1932)
print(var_test)
sd_stats <- df_1932 %>% group_by(treatmentgroup) %>%
summarise(sd = sd(diffpatentswrt1918, na.rm = TRUE))
sd_effects <- sd_stats$sd[2] - sd_stats$sd[1]
effect_reg <- lm(diffpatentswrt1918 ~ treatmentgroup, data = df_1932)
beta_reg <- coef(effect_reg)["treatmentgroup"]
cat("CI: [", beta_reg - 1.96*sd_effects, ",", beta_reg + 1.96*sd_effects, "]")F = 0.77644, p-value = 0.0008145
SD control: 1.772807
SD treated: 2.011905
sd_2 - sd_1 = 0.2390978
beta = 0.6422164
CI: [ 0.1735848 , 1.110848 ]import pandas as pd
import pyfixest as pf
from scipy import stats
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
df_1932 = df[df["year"] == 1932]
g0 = df_1932.loc[df_1932["treatmentgroup"] == 0, "diffpatentswrt1918"].dropna()
g1 = df_1932.loc[df_1932["treatmentgroup"] == 1, "diffpatentswrt1918"].dropna()
sd0, sd1 = g0.std(ddof=1), g1.std(ddof=1)
sd_effects = sd1 - sd0
F_var = sd0**2 / sd1**2
p_two = 2 * min(stats.f.cdf(F_var, len(g0)-1, len(g1)-1),
1 - stats.f.cdf(F_var, len(g0)-1, len(g1)-1))
m7 = pf.feols("diffpatentswrt1918 ~ treatmentgroup", data=df_1932)
beta = m7.coef()["treatmentgroup"]
print(f"F = {F_var:.4f}, p = {p_two:.4f}")
print(f"sd_2 - sd_1 = {sd_effects:.6f}")
print(f"beta = {beta:.6f}")
print(f"CI: [{beta - 1.96*sd_effects:.6f}, {beta + 1.96*sd_effects:.6f}]")SD control: 1.772923
SD treated: 2.011905
sd_2 - sd_1 = 0.238982
F = 0.7765, 2*Pr(F<f) = 0.0008
beta = 0.642216
CI: [0.173812, 1.110620]Result: The treated group has a larger variance (\(\hat{\sigma}_1 = 2.01 > \hat{\sigma}_0 = 1.77\)). The implied SD of treatment effects is \(\approx 0.239\), and the confidence interval for \(\beta_{14}\) using this SD is \([0.174, 1.111]\).
1.10 CQ#10 (cont.): Placebo Test
Perform a placebo test by comparing the variances of
diffpatentswrt1918across all years.
copy "https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.dta" "moser_voena_didtextbook.dta", replace
use "moser_voena_didtextbook.dta", clear
forvalues i = 1900/1939 {
quietly sdtest diffpatentswrt1918 if year==`i', by(treatmentgroup)
display "Year `i': F = " %6.4f r(F) " p = " %6.4f r(p)
}Year 1900: F = 3.2735 p = 0.0000
Year 1901: F = 2.5398 p = 0.0000
Year 1902: F = 3.3703 p = 0.0000
Year 1903: F = 3.1775 p = 0.0000
Year 1904: F = 4.0726 p = 0.0000
Year 1905: F = 3.9685 p = 0.0000
Year 1906: F = 3.5803 p = 0.0000
Year 1907: F = 1.7711 p = 0.0000
Year 1908: F = 2.5510 p = 0.0000
Year 1909: F = 3.7689 p = 0.0000
Year 1910: F = 2.2839 p = 0.0000
Year 1911: F = 1.8151 p = 0.0000
Year 1912: F = 3.7112 p = 0.0000
Year 1913: F = 2.2403 p = 0.0000
Year 1914: F = 2.7455 p = 0.0000
Year 1915: F = 3.6764 p = 0.0000
Year 1916: F = 3.2690 p = 0.0000
Year 1917: F = 1.6222 p = 0.0000
Year 1918: F = . p = .
Year 1919: F = 1.7652 p = 0.0000
Year 1920: F = 1.7161 p = 0.0000
Year 1921: F = 2.6409 p = 0.0000
Year 1922: F = 1.6035 p = 0.0000
Year 1923: F = 1.2684 p = 0.0040
Year 1924: F = 2.3221 p = 0.0000
Year 1925: F = 1.6974 p = 0.0000
Year 1926: F = 2.4032 p = 0.0000
Year 1927: F = 1.5314 p = 0.0000
Year 1928: F = 1.7269 p = 0.0000
Year 1929: F = 1.2982 p = 0.0016
Year 1930: F = 1.1443 p = 0.0999
Year 1931: F = 1.5775 p = 0.0000
Year 1932: F = 0.7764 p = 0.0008
Year 1933: F = 0.8284 p = 0.0134
Year 1934: F = 1.1337 p = 0.1256
Year 1935: F = 0.8164 p = 0.0076
Year 1936: F = 0.7675 p = 0.0005
Year 1937: F = 0.8621 p = 0.0522
Year 1938: F = 0.8692 p = 0.0668
Year 1939: F = 0.7569 p = 0.0002library(haven); library(dplyr)
load(url("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.RData"))
for (yr in 1900:1939) {
df_yr <- df %>% filter(year == yr)
test <- var.test(diffpatentswrt1918 ~ treatmentgroup, data = df_yr)
cat(sprintf("Year %d: F = %.4f, p = %.4f\n", yr, test$statistic, test$p.value))
}Year 1900: F = 3.2735, p = 0.0000
Year 1901: F = 2.5398, p = 0.0000
Year 1902: F = 3.3703, p = 0.0000
Year 1903: F = 3.1775, p = 0.0000
Year 1904: F = 4.0726, p = 0.0000
Year 1905: F = 3.9685, p = 0.0000
Year 1906: F = 3.5803, p = 0.0000
Year 1907: F = 1.7711, p = 0.0000
Year 1908: F = 2.5510, p = 0.0000
Year 1909: F = 3.7689, p = 0.0000
Year 1910: F = 2.2839, p = 0.0000
Year 1911: F = 1.8151, p = 0.0000
Year 1912: F = 3.7112, p = 0.0000
Year 1913: F = 2.2403, p = 0.0000
Year 1914: F = 2.7455, p = 0.0000
Year 1915: F = 3.6764, p = 0.0000
Year 1916: F = 3.2690, p = 0.0000
Year 1917: F = 1.6222, p = 0.0000
Year 1918: F = ., p = .
Year 1919: F = 1.7652, p = 0.0000
Year 1920: F = 1.7161, p = 0.0000
Year 1921: F = 2.6409, p = 0.0000
Year 1922: F = 1.6035, p = 0.0000
Year 1923: F = 1.2684, p = 0.0040
Year 1924: F = 2.3221, p = 0.0000
Year 1925: F = 1.6974, p = 0.0000
Year 1926: F = 2.4032, p = 0.0000
Year 1927: F = 1.5314, p = 0.0000
Year 1928: F = 1.7269, p = 0.0000
Year 1929: F = 1.2982, p = 0.0016
Year 1930: F = 1.1443, p = 0.0999
Year 1931: F = 1.5775, p = 0.0000
Year 1932: F = 0.7764, p = 0.0008
Year 1933: F = 0.8284, p = 0.0134
Year 1934: F = 1.1337, p = 0.1256
Year 1935: F = 0.8164, p = 0.0076
Year 1936: F = 0.7675, p = 0.0005
Year 1937: F = 0.8621, p = 0.0522
Year 1938: F = 0.8692, p = 0.0668
Year 1939: F = 0.7569, p = 0.0002import pandas as pd
from scipy import stats
df = pd.read_parquet("https://raw.githubusercontent.com/anzonyquispe/did_book/main/cc_xd_didtextbook_2025_9_30/Data%20sets/Moser%20and%20Voena%202012/moser_voena_didtextbook.parquet")
for yr in range(1900, 1940):
dfy = df[df["year"] == yr]
g0y = dfy.loc[dfy["treatmentgroup"] == 0, "diffpatentswrt1918"].dropna()
g1y = dfy.loc[dfy["treatmentgroup"] == 1, "diffpatentswrt1918"].dropna()
if len(g0y) > 1 and len(g1y) > 1 and g1y.std(ddof=1) > 0:
F_v = g0y.std(ddof=1)**2 / g1y.std(ddof=1)**2
p_v = 2 * min(stats.f.cdf(F_v, len(g0y)-1, len(g1y)-1),
1 - stats.f.cdf(F_v, len(g0y)-1, len(g1y)-1))
print(f"Year {yr}: F = {F_v:.4f} p = {p_v:.4f}")Year 1900: F = 3.2740 p = 0.0000
Year 1901: F = 2.5396 p = 0.0000
Year 1902: F = 3.3701 p = 0.0000
Year 1903: F = 3.1773 p = 0.0000
Year 1904: F = 4.0723 p = 0.0000
Year 1905: F = 3.9683 p = 0.0000
Year 1906: F = 3.5801 p = 0.0000
Year 1907: F = 1.7712 p = 0.0000
Year 1908: F = 2.5509 p = 0.0000
Year 1909: F = 3.7687 p = 0.0000
Year 1910: F = 2.2838 p = 0.0000
Year 1911: F = 1.8153 p = 0.0000
Year 1912: F = 3.7110 p = 0.0000
Year 1913: F = 2.2402 p = 0.0000
Year 1914: F = 2.7453 p = 0.0000
Year 1915: F = 3.6762 p = 0.0000
Year 1916: F = 3.2688 p = 0.0000
Year 1917: F = 1.6223 p = 0.0000
Year 1918: (undefined - reference year)
Year 1919: F = 1.7651 p = 0.0000
Year 1920: F = 1.7159 p = 0.0000
Year 1921: F = 2.6407 p = 0.0000
Year 1922: F = 1.6034 p = 0.0000
Year 1923: F = 1.2684 p = 0.0040
Year 1924: F = 2.3220 p = 0.0000
Year 1925: F = 1.6973 p = 0.0000
Year 1926: F = 2.4031 p = 0.0000
Year 1927: F = 1.5313 p = 0.0000
Year 1928: F = 1.7268 p = 0.0000
Year 1929: F = 1.2981 p = 0.0016
Year 1930: F = 1.1442 p = 0.1002
Year 1931: F = 1.5777 p = 0.0000
Year 1932: F = 0.7765 p = 0.0008
Year 1933: F = 0.8284 p = 0.0134
Year 1934: F = 1.1336 p = 0.1258
Year 1935: F = 0.8164 p = 0.0076
Year 1936: F = 0.7675 p = 0.0005
Year 1937: F = 0.8621 p = 0.0522
Year 1938: F = 0.8692 p = 0.0668
Year 1939: F = 0.7569 p = 0.0002Result: Pre-TWEA years show \(F > 1\) (control group has higher variance), while post-1930 years show \(F < 1\) (treated group variance increases). The pattern is consistent across all three languages.